The landscape of oncology has been fundamentally reshaped by the advent of adoptive cell therapies, particularly T cell receptor-engineered T cell (TCR-T) therapy. Unlike Chimeric Antigen Receptor (CAR) T cells, which are restricted to surface antigens, TCRs can recognize intracellular targets presented as peptides on the Major Histocompatibility Complex (MHC). This unlocks a vast repository of tumor-associated antigens (TAAs) and tumor-specific neoantigens for therapeutic intervention.
However, discovering and optimizing a naturally occurring TCR for clinical use is a remarkably slow and arduous process. Nature’s repertoire, while vast, is constrained by thymic selection, often resulting in low-affinity TCRs against self-antigens. To harness the full potential of TCR-T therapies, the pharmaceutical industry is increasingly turning to artificial intelligence. AI TCR design has emerged as a transformative approach, allowing researchers to engineer synthetic TCRs that are fine-tuned for therapeutic efficacy.
Despite the immense promise of computational modeling, engineering an effective TCR is an intricate balancing act. In this comprehensive guide, we will explore the core pillars of AI-driven TCR design for oncology, focusing on the three most critical constraints: specificity, TCR cross-reactivity, and developability.
The Paradigm Shift: From Discovery to AI TCR Design
Historically, TCR discovery relied on isolating T cells from patients or humanized mice, sequencing the receptors, and validating them through exhaustive in vitro assays. This empirical approach is time-consuming, costly, and heavily dependent on chance.
AI shifts this paradigm from “discovery” to “design.” By leveraging deep learning, protein language models, and advanced structural prediction algorithms (such as AlphaFold-based multimer predictions), scientists can now map the vast TCR-pMHC (peptide-MHC) sequence space. AI models can learn the complex structural grammar of the immunological synapse, enabling the de novo generation of TCR sequences or the targeted affinity maturation of existing ones.
However, designing a TCR in silico is not just about maximizing binding affinity to the target tumor antigen. An over-optimized TCR can be catastrophic if it fails to adhere to the strict biological constraints required for safety and manufacturability.
Constraint 1: Maximizing Target Specificity
Specificity is the foundation of any targeted oncological therapy. The TCR must recognize its cognate peptide-MHC complex with high precision to effectively trigger T cell activation and subsequent tumor lysis.
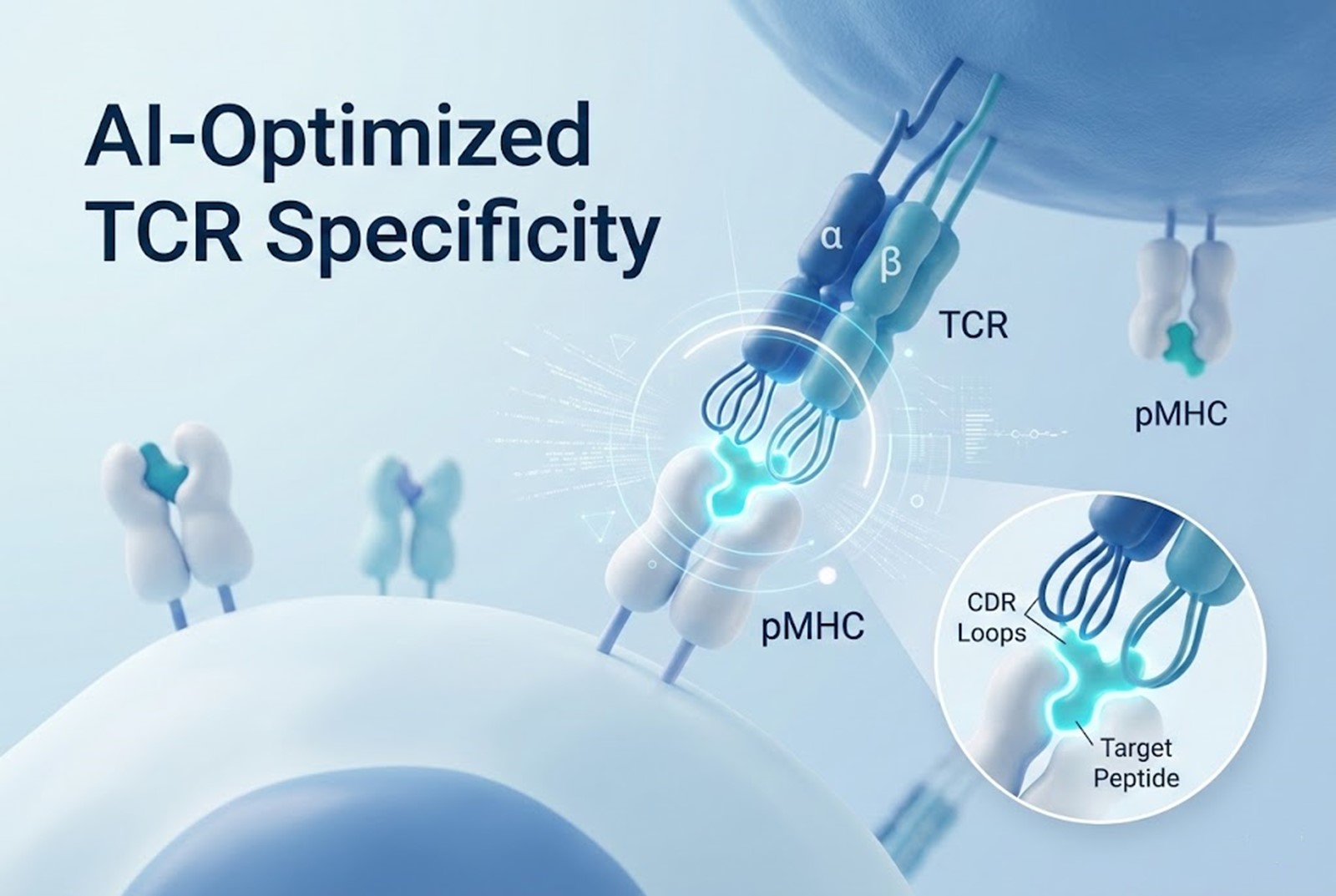
The Challenge of Tumor Antigens
In oncology, many targets are TAAs—proteins that are overexpressed in cancer cells but still present at low levels in healthy tissues (e.g., NY-ESO-1, MAGE-A4). Natural TCRs against these antigens typically possess low affinity (Kd in the micromolar range) to prevent autoimmunity. While AI can easily suggest mutations in the Complementarity-Determining Regions (CDRs)—particularly the highly variable CDR3 loop—to boost affinity into the nanomolar or even picomolar range, this must be done with extreme precision.
AI-Driven Specificity Optimization
Modern AI TCR design algorithms evaluate specificity by modeling the 3D conformation of the TCR-pMHC interface. Machine learning models analyze:
-
Hydrogen bonding networks: Ensuring stable interactions between the TCR $\alpha$ and $\beta$ chains and the target peptide.
-
Shape complementarity: Calculating the geometric fit between the TCR and the unique topography of the MHC-presented peptide.
-
Energetic landscapes: Utilizing molecular dynamics (MD) simulations integrated with neural networks to predict the free energy of binding ΔG.
By computing these parameters, AI ensures that the enhanced affinity is driven strictly by interactions with the peptide rather than non-specific binding to the MHC backbone, preserving the delicate balance required for true target specificity.
Constraint 2: Mitigating TCR Cross-Reactivity
While specificity ensures the TCR hits its intended target, evaluating TCR cross-reactivity is essential to ensure it does not hit unintended targets. In the context of engineered T cell therapies, off-target toxicity can be fatal. If a high-affinity engineered TCR recognizes a structurally similar, but entirely different peptide presented on healthy tissues (a phenomenon known as molecular mimicry), it can lead to severe adverse events.
The Complexity of the Human Peptidome
The human body presents hundreds of thousands of unique peptides on HLA molecules. Screening a novel TCR against the entire healthy human peptidome in vitro is virtually impossible. This is where AI becomes indispensable.
Predicting and Preventing Off-Target Effects
Managing TCR cross-reactivity is arguably the most challenging computational constraint. AI platforms tackle this through large-scale predictive screening:
-
Sequence-Based Motif Scanning: Deep learning algorithms scan the human proteome to identify peptides with sequence homology to the target tumor antigen. The AI scores these sequences based on their likelihood of being processed and presented by the specific HLA allele.
-
Structural Cross-Reactivity Modeling: Because TCRs can bind peptides with low sequence identity but similar 3D conformations, advanced structural AI models predict the binding affinity of the engineered TCR against a library of “decoy” pMHC complexes.
-
Negative Selection Algorithms: During the AI TCR design loop, models are penalized for mutations that increase the predicted binding score to known healthy tissue antigens. This in silico “thymic selection” filters out dangerous candidates long before they reach the laboratory.
By rigorously profiling TCR cross-reactivity computationally, researchers can de-risk therapeutic candidates early in the pipeline, saving millions of dollars and preventing clinical trial failures.
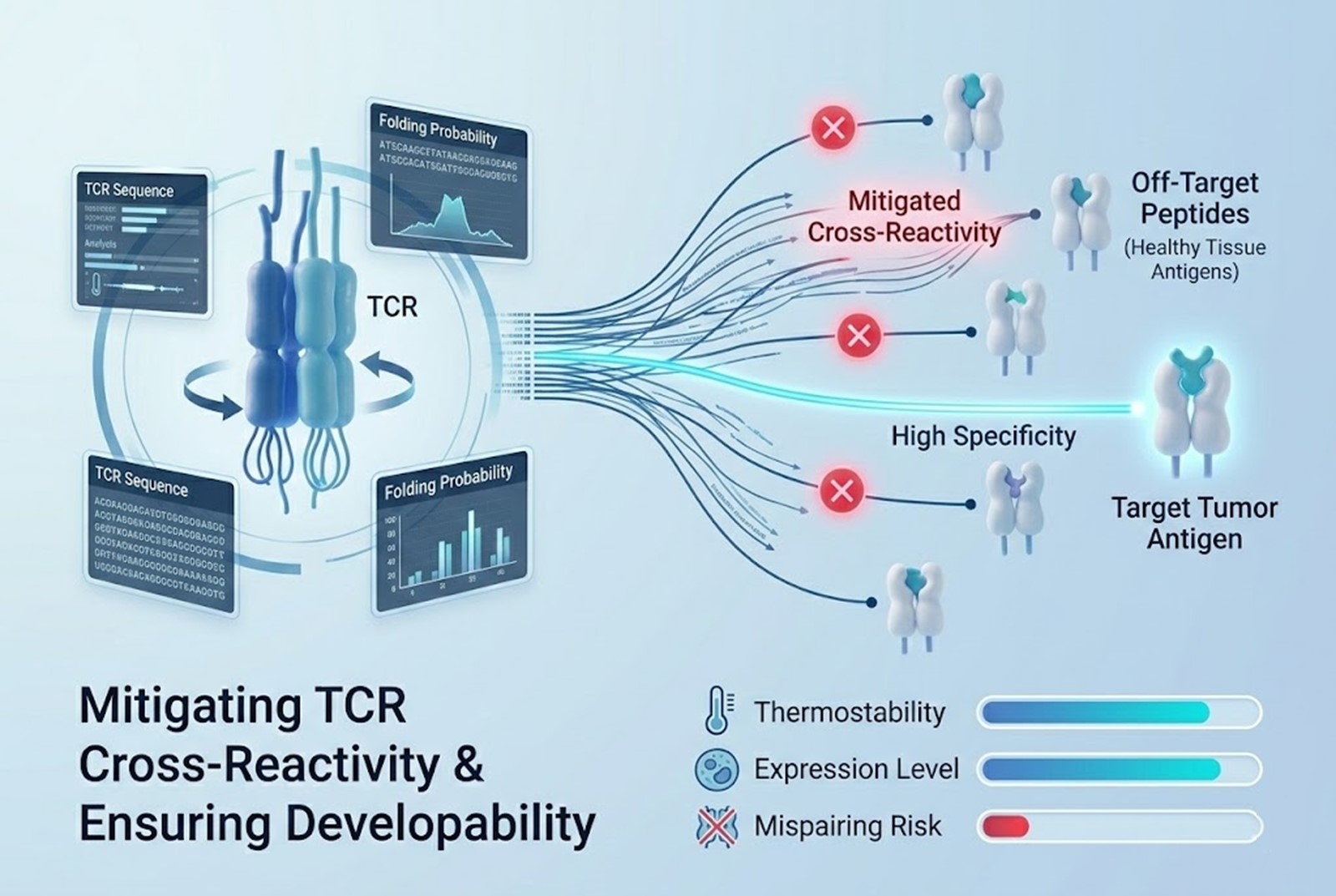
Constraint 3: Ensuring Clinical Developability
A TCR may exhibit flawless specificity and zero predicted cross-reactivity in silico, but it is useless if it cannot be manufactured or expressed effectively in a patient’s T cells. Developability refers to the biophysical properties that make a protein viable as a therapeutic product.
Overcoming Biophysical Hurdles
When AI mutates a natural TCR sequence to enhance affinity, it often inadvertently introduces biophysical liabilities. Key developability constraints include:
-
Thermostability: The engineered TCR must fold correctly and remain stable at physiological temperatures. Unstable TCRs degrade quickly, reducing the persistence and efficacy of the T cell therapy.
-
Expression Levels: The synthetic TCR must traffic efficiently to the T cell surface. Poor expression limits the avidity of the T cell for the tumor.
-
Mispairing: In traditional TCR-T therapy, the engineered $\alpha$ and $\beta$ chains can mispair with the patient’s endogenous TCR chains. This not only reduces the number of functional therapeutic TCRs on the cell surface but can also create novel, unpredictable TCRs with unknown and potentially dangerous cross-reactivities.
AI for Developability Optimization
Advanced AI pipelines integrate developability predictions directly into the generative design phase:
-
Protein Language Models (pLMs): Models trained on millions of protein sequences can evaluate the “naturalness” of a designed TCR sequence. Mutations that lower the naturalness score often correlate with poor folding and stability, allowing the AI to discard them.
-
Structural Stability Metrics: AI predicts the structural impact of amino acid substitutions, avoiding mutations that expose hydrophobic patches (which lead to aggregation) or disrupt essential stabilizing bonds.
-
Framework Engineering: AI can design modifications outside the CDR loops, such as introducing artificial disulfide bonds between the $\alpha$ and $\beta$ constant domains or utilizing murine constant regions, to enhance stability and prevent endogenous mispairing.
The Future: A Multi-Objective AI Design Workflow
The true power of AI in oncology lies in its ability to perform multi-objective optimization. Instead of optimizing affinity, checking for cross-reactivity, and then testing for developability in isolated, sequential steps, modern AI frameworks handle these constraints simultaneously.
By defining the ideal therapeutic profile—high affinity for the tumor neoantigen, strict negative selection against the healthy peptidome, and high thermostability—researchers can use generative AI to output a curated library of TCR candidates that satisfy all criteria at once.
As machine learning models ingest more high-quality, high-throughput wet-lab data regarding TCR-pMHC interactions, the fidelity of these predictions will only increase. This tight integration of computational design and biological validation is accelerating the timeline from tumor biopsy to personalized, safe, and highly effective cell therapies.
Accelerate Your Pipeline with Creative Biolabs
At Creative Biolabs, we understand that navigating the complexities of computational immunology requires cutting-edge tools and deep domain expertise. We offer a comprehensive suite of AI-driven solutions tailored to accelerate your oncology therapeutics pipeline, from early-stage discovery to clinical developability optimization.
Our specialized services can help you overcome developmental bottlenecks and bring safer, more effective treatments to market faster:
-
AI TCR Design Service: Discover and engineer high-affinity, highly specific T cell receptors with our advanced predictive models.
-
AI Gene & Cell Therapy Development Service: Leverage artificial intelligence to streamline the design and optimization of advanced cell-based therapies.
-
AI Multi-specific Antibody Design Service: Develop complex multi-specific biologics with optimized binding interfaces and reduced structural liabilities.
-
AI Soluble Receptor Design Service: Engineer stable, high-affinity soluble receptors for targeted therapeutic applications.
-
AI Oncology mRNA Vaccine Development Service: Utilize computational tools to design highly immunogenic and stable mRNA constructs for cancer immunotherapy.
-
AI Antibody Developability Optimization Platform: Ensure your biologic candidates possess the optimal biophysical properties for successful clinical translation and manufacturing.
