The landscape of targeted therapeutics and diagnostics is experiencing a profound paradigm shift. For decades, monoclonal antibodies have dominated the field of targeted biologicals. However, the rise of oligonucleotides has brought a formidable contender to the forefront: aptamers. Often referred to as “chemical antibodies,” these single-stranded DNA or RNA molecules can fold into complex three-dimensional structures, allowing them to bind to specific targets with high affinity and specificity.
Despite their immense potential, the traditional path to discovering viable aptamers has historically been fraught with bottlenecks, primarily relying on highly repetitive, time-consuming laboratory processes. Today, the integration of artificial intelligence is fundamentally rewriting this narrative. AI aptamer design is rapidly transforming the field, offering unprecedented speed and precision. In this comprehensive guide, we will explore how advanced computational algorithms are enabling faster iteration, moving seamlessly from early motif discovery to rigorous candidate screening.
What Aptamers Solve in Modern Therapeutics and Diagnostics
Before delving into the mechanics of AI-driven discovery, it is crucial to understand why aptamers are garnering such intense interest from pharmaceutical developers and researchers. While antibodies are excellent tools, they have inherent limitations that aptamers are perfectly positioned to solve.
-
Superior Tissue Penetration: Aptamers are substantially smaller than monoclonal antibodies (typically 10-30 kDa compared to 150 kDa for IgGs). This compact size allows them to penetrate dense tissues, such as solid tumors, much more effectively, improving the delivery of conjugated payloads or imaging agents.
-
Low Immunogenicity and Toxicity: Because they are nucleic acids, aptamers generally evade the host immune system, minimizing the risk of adverse immune responses—a common challenge in biologic therapies.
-
Unmatched Thermal and Chemical Stability: Unlike proteins that denature irreversibly under heat or chemical stress, DNA and RNA aptamers can often return to their functional folded state after the stressor is removed. This grants them a significantly longer shelf life and eliminates the strict cold-chain requirements associated with many biologics.
-
Efficient Chemical Synthesis: Aptamers are synthesized chemically rather than biologically. This means batch-to-batch variation is virtually non-existent, production scales up predictably, and site-specific modifications (like adding fluorophores, PEGylation, or drug conjugates) are straightforward and highly controlled.
Despite these advantages, the primary barrier to widespread aptamer adoption has been the unpredictable and laborious discovery process.
The Evolution of Discovery: SELEX vs AI
For over thirty years, the gold standard for finding target-specific aptamers has been the SELEX (Systematic Evolution of Ligands by Exponential Enrichment) process.
The Traditional SELEX Bottleneck
SELEX begins with a massive, randomized library of oligonucleotides (often 1013 to 1015 sequences). This library is incubated with the target molecule. Sequences that do not bind are washed away, while the bound sequences are eluted and amplified using PCR. This cycle is typically repeated 10 to 15 times to enrich the pool with high-affinity binders.
While conceptually elegant, SELEX is notoriously tedious. It can take months to complete, is prone to PCR bias (where easily amplifiable sequences outcompete actual high-affinity binders), and frequently fails when targets are complex or unstable. Furthermore, SELEX only explores a minuscule fraction of the theoretical sequence space; even a massive library of 1015 molecules barely scratches the surface of the possible combinations for a 40-nucleotide sequence (which is 440, or roughly 1.2×1024.
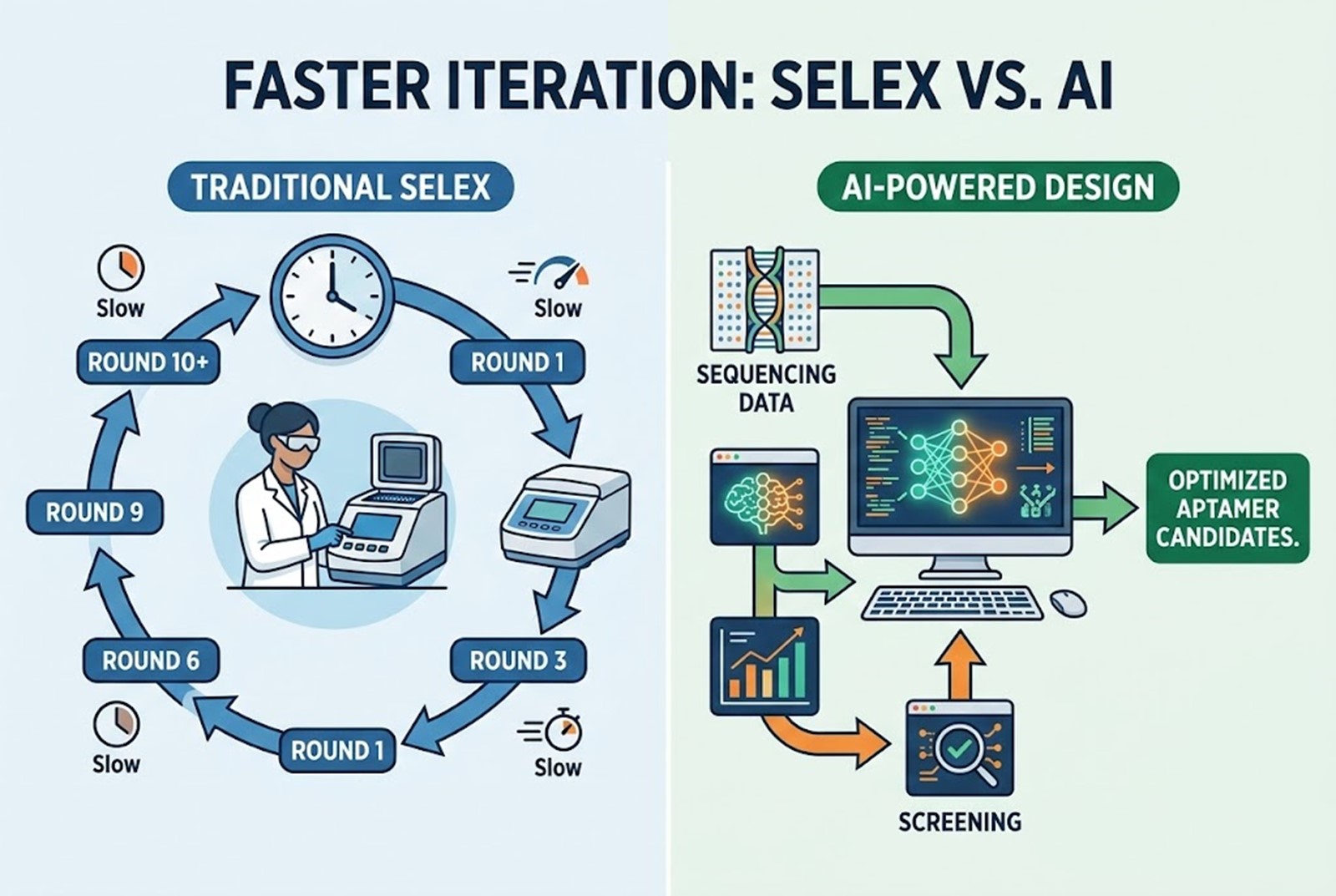
The AI Advantage
AI aptamer design directly addresses the pitfalls of SELEX. By integrating high-throughput sequencing (HTS) data with deep learning architectures, AI bypasses the need for endless physical iterations.
Instead of relying solely on physical enrichment, AI algorithms—such as Recurrent Neural Networks (RNNs), Transformers, and Graph Neural Networks (GNNs)—analyze the sequencing data from just the first few rounds of a SELEX experiment. The AI learns the underlying sequence-to-function and sequence-to-structure relationships. It can then predict the binding affinities of sequences that were never even physically synthesized, effectively expanding the search space exponentially and generating de novo sequences with optimized properties.
Motif Discovery: The Core of High-Affinity Binding
The true power of AI in this domain begins with motif discovery. An aptamer’s ability to bind to its target is dictated by its three-dimensional conformation, which is, in turn, driven by specific secondary structural motifs—such as stem-loops, hairpins, bulges, pseudoknots, and G-quadruplexes.
In traditional bioinformatics, finding these conserved motifs across millions of sequencing reads is a computationally heavy, statistically noisy endeavor. AI models excel at pattern recognition in noisy datasets. Unsupervised learning algorithms and variational autoencoders (VAEs) can cluster vast datasets of oligonucleotide sequences, identifying hidden, non-obvious structural and sequence motifs that correlate with binding affinity.
By mapping the “grammar” of these motifs, AI systems can pinpoint exactly which localized regions of the aptamer are interacting with the target’s active sites. This allows for the intelligent truncation of long sequences, cutting away unnecessary nucleotides to create minimized, highly potent aptamers that are cheaper to synthesize and more stable in vivo.
Candidate Triage and In Silico Aptamer Screening
Once a vast pool of potential sequences has been generated or enriched by the AI, the next critical phase is triage. This is where aptamer screening becomes a purely computational, high-throughput endeavor before any wet-lab resources are expended.
Predictive Scoring and Molecular Docking
Using advanced deep learning models, candidates are subjected to rigorous in silico aptamer screening. Algorithms predict the secondary and tertiary structures of the RNA or DNA sequences. Concurrently, tools utilizing physics-based modeling and machine learning predict how these 3D structures will interact with the target protein.
Molecular docking simulations generate millions of potential binding poses. Sophisticated scoring functions evaluate the binding free energy (ΔG), hydrogen bonding, electrostatic interactions, and Van der Waals forces.
Multi-Parameter Optimization
Crucially, AI does not just screen for affinity. A successful therapeutic candidate must also possess favorable pharmacokinetic properties. During the triage phase, AI models simultaneously evaluate and filter candidates based on:
-
Specificity: Predicting potential cross-reactivity or off-target binding by screening the aptamer against a database of non-target human proteins.
-
Stability: Estimating the thermodynamic stability and nuclease resistance of the predicted secondary structures.
-
Synthesizability: Ensuring the sequence does not form pathological aggregations that complicate chemical manufacturing.
This multi-objective optimization ensures that only the most robust, drug-like candidates survive the triage process.

Wet-Lab Validation: Bridging the Computational and Physical Worlds
It is a misconception that AI completely replaces the laboratory. Rather, AI transforms the laboratory from a place of blind searching into a site of targeted confirmation.
After in silico aptamer screening reduces the candidate pool from billions to a highly curated list of 10 to 50 top-tier sequences, these select few are physically synthesized. Wet-lab validation then confirms the computational predictions.
Scientists employ high-precision biophysical techniques to validate these candidates. Surface Plasmon Resonance (SPR) and Bio-Layer Interferometry (BLI) are used to measure real-time binding kinetics, yielding precise association Ka and dissociation Kb rates. Flow cytometry and confocal microscopy may be utilized to verify binding in complex cellular environments if the target is a cell-surface receptor. Because the AI has already filtered out the “noise,” the success rate in this final wet-lab validation phase is exponentially higher than in traditional discovery campaigns.
Deliverables in AI-Driven Aptamer Discovery
When researchers leverage an AI-driven platform for aptamer discovery, the output is comprehensive and actionable. The deliverables go far beyond a simple list of A, T, C, and Gs.
Clients typically receive:
-
A Ranked Candidate List: The top computationally and physically validated aptamer sequences.
-
Detailed Binding Kinetics: Comprehensive Kb values and affinity profiles derived from wet-lab validation.
-
Structural Modeling: 3D in silico models illustrating the predicted binding interface and molecular interactions between the aptamer and the target.
-
Motif Analysis: Documentation of the critical structural motifs driving the binding interaction, providing a foundation for future optimization or truncation.
-
Specificity Profiles: Data detailing the aptamer’s behavior against negative control targets to guarantee specific binding.
By providing this wealth of data, AI empowers researchers to confidently advance their programs into preclinical testing at a fraction of the traditional cost and time.
Accelerate Your Pipeline with Creative Biolabs
At Creative Biolabs, we leverage cutting-edge artificial intelligence to revolutionize the discovery and development of next-generation therapeutics. Whether you are developing novel diagnostics, targeted drug delivery systems, or complex biologics, our AI-powered platforms are designed to accelerate your workflow, reduce costs, and mitigate clinical risks.
Explore our specialized AI-driven services below to see how we can support your next breakthrough:
-
AI Aptamer Design Service: Rapidly discover high-affinity, high-specificity aptamers utilizing our advanced generative AI and in silico screening platforms.
-
AI Gene & Cell Therapy Development Service: Optimize vector design, enhance payload delivery, and predict off-target effects in your gene and cell therapy pipelines.
-
AI Multi-Specific Antibody Design Service: Engineer complex bispecific and multi-specific antibodies with optimized binding interfaces and superior developability.
-
AI TCR Design Service: Accelerate your adoptive cell therapy research with AI-guided T-cell receptor discovery and affinity maturation.
-
AI Oncology mRNA Vaccine Development Service: Accelerate cancer vaccine development with AI-driven neoantigen prediction and mRNA sequence optimization for maximum immunogenicity.
